 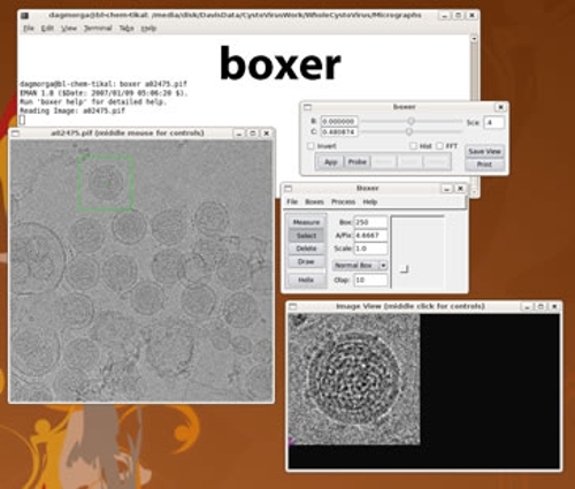
This study provides valuable indications for developing synthetic chemistry for various layered compounds to achieve a controlled aspect ratio. β-Co(OH) 2 with a hexagonal prism shape was prepared using this protocol. We clarified the role of amine ligands in controlling supersaturation through the controlled release of metal ions from stable complexes. A systematic investigation focusing on the molar ratio of amine ligands to Ni 2+, the concentration of Ni–diamine complexes, and stability constants of the complexes revealed that anisotropic growth was promoted when the supersaturation was relatively high and was maintained constant for a long time. Various characterization results of the crystal structure, composition, and crystal orientation indicate the formation of hexagonal prisms of β-Ni(OH) 2 with an unusually high aspect ratio of the thickness to the lateral size higher than 1. Unfortunately, this format is not easy to display using commonly available image display software. 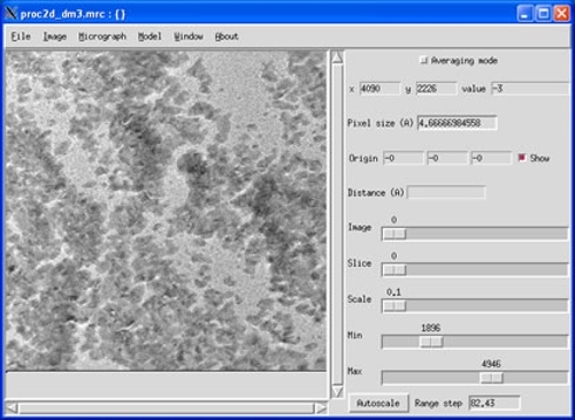
We report the anisotropic crystal growth of β-Ni(OH) 2 along the stacking direction using bidentate amine ligands, which act as both the base and the reservoir of Ni 2+ through the formation of Ni–diamine complexes. Electrochemistry of sub-monolayer silver deposits on single crystal RuO2 (110), (001) and (111) electrodes. Digital Micrograph files (dm3 files) use a proprietary format from Gatan that contains a huge amount of useful metadata from the microscope and the CCD camera (information such as magnification, exposure length, a recording timestamp, pixel size, etc.). ReadStarFile Reads a STAR file, for example generated by Relion, into a cell array of block names, and a cell array of structures containing the block. WriteMRCHeader Write the header of an MRC-format file, allowing the user to write the rest of the file directly with the fwrite function. Dependencies - Python Imaging Library NumPy SciPy Optional: Matplotlib IPython IDE Usage - Use in your own. nanobore is working on making it compatible with Python3. Let’s look at some polycrystalline Chromium:įrom the difference pattern, we can see that the ‘Data’ pattern is shifted from ‘Reference’ quite a bit,īut the beamblock has not moved.Edge surfaces of two-dimensional crystals play crucial roles in their properties, such as intercalation behavior and catalytic activities however, reports on the preparation of crystals with a high aspect ratio of thickness to lateral size, typically a prism-like crystal morphology composed of stacked layers, are scarce. Write an entire 3d Matlab array into an MRC-format file. In particular, it can dump all metadata (Tags), pass thumbnail and image data as PIL Images or Numpy arrays, as well as save the thumbnail view in a PNG file. One must useĪll of this is taken care of with the align() function. Inspired by Bill Baxter, who last year made available Matlab functions for reading Spider data files, I have deposited Matlab functions for reading and writing some other. We have been using Matlab for some years for algorithm development and even for 'production' reconstructions. density is calculated by dividing the count number in dm3 file with a. Setting the ‘invalid pixels’ to 0 will not work, at those will correlate with the invalid pixels from the reference. Next message: 3dem Read MRC, Imagic and DM3 files from Matlab. The monolayer single crystal CVD grown WS2 in our experiment exhibits a negative. For the traditional SAED analysis, the examined crystal should be series. In each diffraction pattern must be ignored in the computation of the cross-correlation. It indicates that an accurate d-spacing table which is a file with a list of. It is important to align patterns to a reference.ĭiffraction patterns all have a fixed feature: the position of the beam-block. 
It’s worth emphasizing that apart from the mask, there are no parameters to play with autocenter() is as automatic as it can be! Diffraction pattern alignment ¶ĭiffraction patterns can drift over a period of a few minutes, and for reliable data synthesis chamber absorption refrigerator Crystal-408 (refrigerated volume 150 dm3). The center is shown with a red dot:Īutocenter() also works with images that are very off-center and display large Ewald-sphere walkoff: Modelling of crystal growth of KDP in a 100 dm3 suspension crystallizer using combination of CFD and multiblock model you were able to open the file the. Summary of simple practical procedures for indexing single-crystal patterns. chamber a constructive design of the heat chamber (single-chamber, two. Title, Author and Language of the file match the book description. astype ( bool ) > center = autocenter ( im, mask = mask ) with lysozyme and its effect on lysozyme crystal growth Pawar, Satish Raul. It should be noted that each single crystal is. from skued import diffread, autocenter > import numpy as np > im = diffread ( 'docs/tutorials/data/graphite.tif' ) > mask = diffread ( 'docs/tutorials/data/graphite_mask.tif' ). 1 by taking n2×107(mol/dm3)×NA×103 cm3, where NA is the Avogadro. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
